Article courtesy of Stephen Sautner, Wildlife Conservation Society
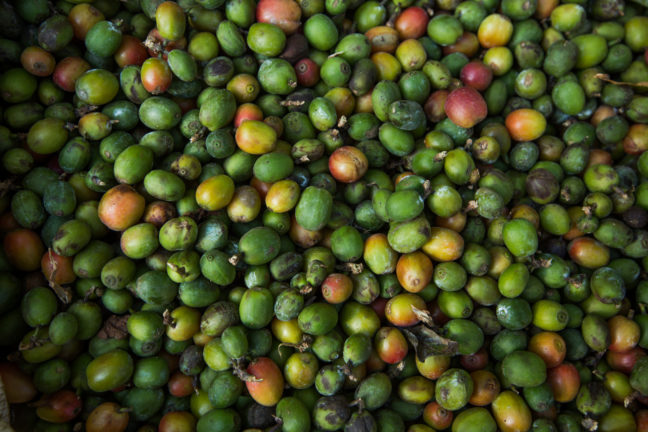
It’s an age-old debate for coffee lovers. Which is better: Arabica beans with their sweeter, softer taste, or the bold, deep flavor of Robusta beans? A new study by the Wildlife Conservation Society (WCS), Princeton University, and the University of Wisconsin-Madison appearing in the journal Scientific Reports has taken the question to unlikely coffee aficionados: birds.

The researchers, led by WCS Associate Conservation Scientist Krithi Karanth, surveyed bird diversity in coffee agroforests in India’s Western Ghats region. Previous research has demonstrated that shade-grown coffee (typically Arabica) can harbor substantial levels of biodiversity. But coffee production is globally shifting toward Robusta, which uses a more intensive full-sun agricultural systems, which may pose deleterious impacts for forest wildlife.
What the researchers found was surprising: although Arabica avian assemblages were more species rich, Robusta nevertheless offered substantial biodiversity benefits, and supported higher densities of several sensitive avian populations such as frugivores, birds that primarily eat fruit. In addition, farmers use less pesticides in the more disease-resistant Robusta farmlands.
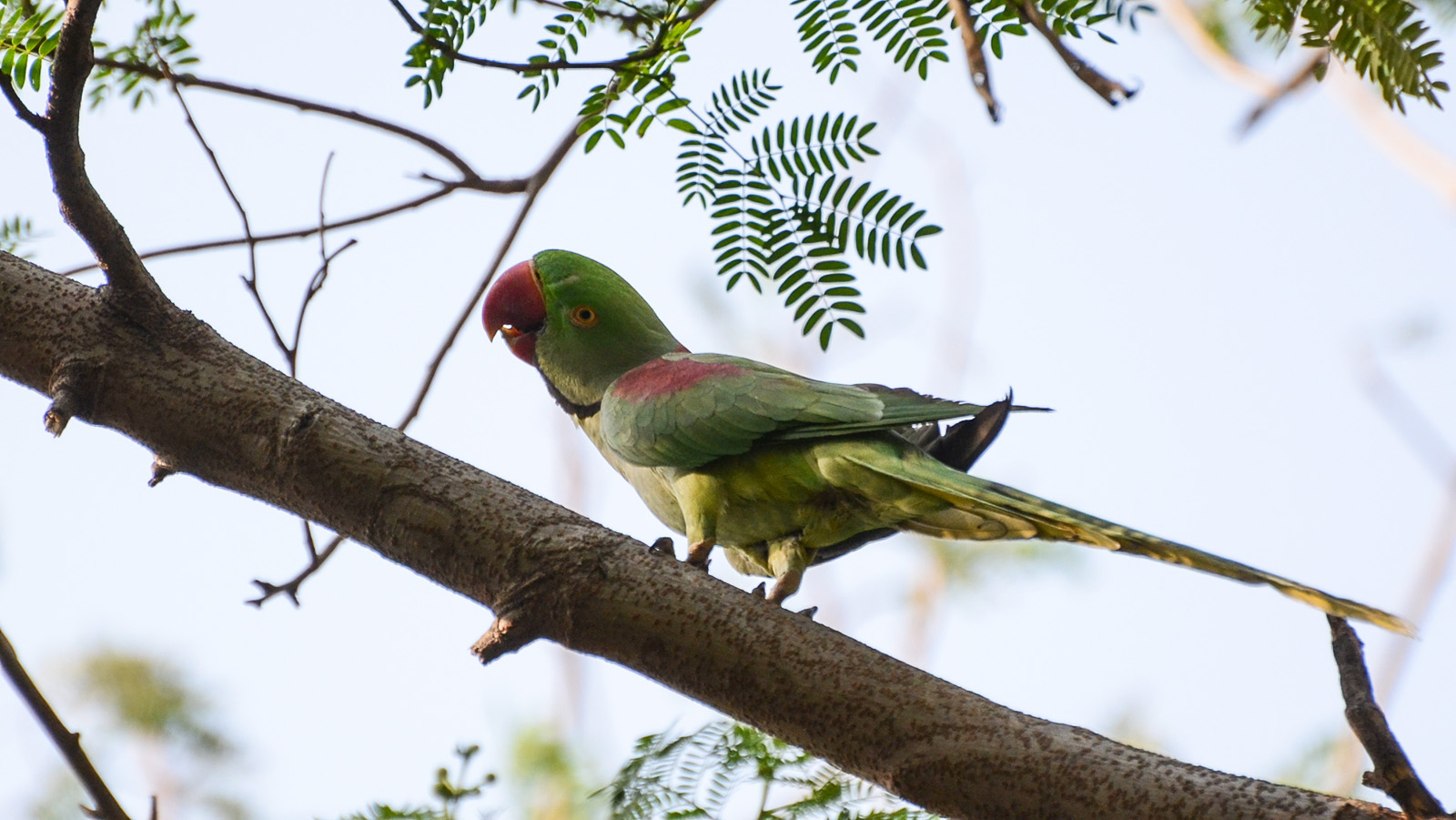
The authors found a total of 79 forest-dependent species living in the coffee plantations they surveyed, including three species listed by the International Union for Conservation of Nature as threatened: Alexandrine parakeet (Psittacula eupatria), grey-headed bulbul (Pycnonotus priocephalus) and the Nilgiri wood pigeon (Columba elphinstonii). Plantations can harbor a diversity of mammals, amphibians and tree species, too.
The study has important implications as coffee production is an increasingly important driver of landscape transformation, and shifts between different coffee bean species are a major dimension of agroforestry trends. The authors say that coffee certification efforts should prioritize maintaining native canopy shade trees to ensure that coffee landscapes can continue providing biodiversity benefits.
Said Dr. Karanth, “Coffee farms already play a complementary role to protected areas in a country like India where less than four percent of land is formally protected. Therefore, building partnerships with largely private individual and corporate land holders will provide much needed safe-passage and additional habitats for birds and other species.”
Indian robusta has relatively high “cup scores” (flavor ratings) by coffee experts, is disease-resistant, and commands a price premium.
Said lead author Charlotte Chang, who analyzed the data while a graduate student in the Department of Ecology and Evolutionary Biology at Princeton University and is now a postdoctoral researcher at the National Institute for Mathematical and Biological Synthesis at the University of Tennessee, Knoxville: “An encouraging result of the study is that coffee production in the Western Ghats, a global biodiversity hotspot, can be a win-win for birds and farmers.”
Chang’s advisers at Princeton were Simon Levin,the James S. McDonnell Distinguished University Professor in Ecology and Evolutionary Biology, and David Wilcove, professor of ecology and evolutionary biology and public affairs and the Princeton Environmental Institute. The senior author on the paper is Paul Robbins, director of the Nelson Institute for Environmental Studies at the University of Wisconsin-Madison.
Funding for this project was provided by the National Science Foundation (grant number 1265223 and a Graduate Research Fellowship), Oracle, and Graduate Research Opportunities Worldwide with USAID.
The study, “Birds and beans: Comparing avian richness and endemism in arabica and robusta agroforests in India’s Western Ghats,” by Charlotte H. Chang, Krithi K. Karanth and Paul Robbins, was published in the journal Scientific Reports online on Feb. 16, 2018. DOI 10.1038/s41598-018-21401-1.










You must be logged in to post a comment.