
By Catherine Zandonella, Office of the Dean for Research
A group of prominent researchers from seven institutions including Princeton University are calling for the establishment of a worldwide program to collect and test blood and other human bodily fluids to aid in the study and prevention of emerging infectious diseases such as the mosquito-borne Zika fever, which is caused by the Zika virus and has spread throughout Latin America and the Caribbean since early 2015.
In an article published in The Lancet April 5, the authors call for the creation of a World Serology Bank that would include storage repositories located around the world that use a common set of best practices to collect and evaluate blood and other bodily fluids from people infected by various pathogens. The resulting data would help scientists better understand the susceptibility of humans to emerging diseases such as Zika fever. The information could be shared widely among scientists who track disease.
The authors include Princeton’s C. Jessica Metcalf, an assistant professor of Ecology and Evolutionary Biology and Public Affairs in the Woodrow Wilson School for Public and International Affairs, and Bryan Grenfell, the Kathryn Briger and Sarah Fenton Professor of Ecology and Evolutionary Biology and Public Affairs. In the interview below, Metcalf explains more about the World Serology Bank proposal.
Why is it important to create a World Serology Bank?
Serology is the study of bodily fluids including serum, the part of the blood that contains antibodies, with the aim of detecting the body’s immune response to pathogens. Serology provides us with the clearest window we have onto the landscape of susceptibility to pathogens across human populations, and the consequent risk of outbreaks. A World Serology Bank would shed light on the global risk of infectious disease outbreaks, and would be of tremendous public health benefit.
Why do you feel it is important to address this issue now?
With the emergence of pathogens like Zika virus, it becomes ever more important to understand what enhances or limits the global spread of these pathogens, and what the consequences of such spread may be across our pathogen communities. A World Serology Bank would provide a powerful mechanism toward such a global perspective.
What are the challenges involved in creating the Bank?
Challenges range from developing systems for collecting fluids, which can be done on a regular schedule or during specific disease events, to methods for sera storage and sera testing. Other challenges include defining who will administer the World Serology Bank, and what global data-sharing agreements will be put in place. Finally, we will need to develop new methods to translate what we learn from the evaluation of sera, such as patterns of susceptibility to specific pathogens, or protection from those pathogens. These methods will be driven by the underlying biology, and are likely to require an array of analytical innovations.
The article, “Use of serological surveys to generate key insights into the changing global landscape of infectious disease,” by C. Jessica E Metcalf, Jeremy Farrar, Felicity T. Cutts, Nicole E. Basta, Andrea L. Graham, Justin Lessler, Neil M. Ferguson, Donald S. Burke and Bryan T. Grenfell was published online in the journal The Lancet on April 5, 2016. http://dx.doi.org/10.1016/S0140-6736(16)30164-7.

 One of the world’s deadliest mosquitoes sustains its taste for human blood thanks in part to a genetic tweak that makes it more sensitive to human odor, according to new research.
One of the world’s deadliest mosquitoes sustains its taste for human blood thanks in part to a genetic tweak that makes it more sensitive to human odor, according to new research.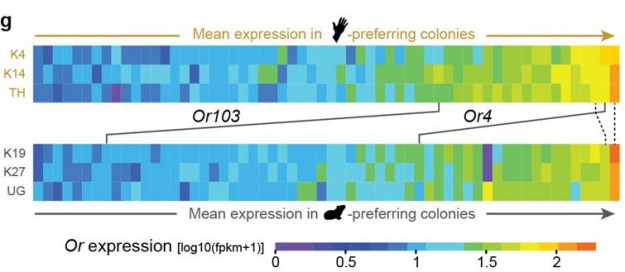
You must be logged in to post a comment.